Non-GMO Testing Update
Results from the Envirologix TotalTrait GMO Soy comb will now include a total GMO estimate value in the final report. This value is the total trait level of the four proteins evaluated in the test: CP4 EPSPS (RoundupReady), PAT/pat (LibertyLink), DMO (Dicamba), and 2mEPSPS (Balance GT/27).
The Envirologix kit has shown to produce false-positive results with the Liberty traits. The Liberty trait expressed in XtendFlex varieties differs from the GT27 varieties. When these two trait stacks are in the same sample, the detection level will generally appear to be much higher than either the dicamba or GT27. This will give the appearance of a higher adventitious presence (AP) level when in reality, the detection of AP as a whole is limited by that of the dicamba and GT27.
In a recent update, Envirologix added a “TotalTrait GMO Soy Sum” to the reader’s software. The percentages of dicamba and GT/27 are compared with the Liberty detected, and the final total is normalized across those traits to produce a more accurate non-GMO or AP total. This value will be added to the report, below the individual protein percentages as shown below.

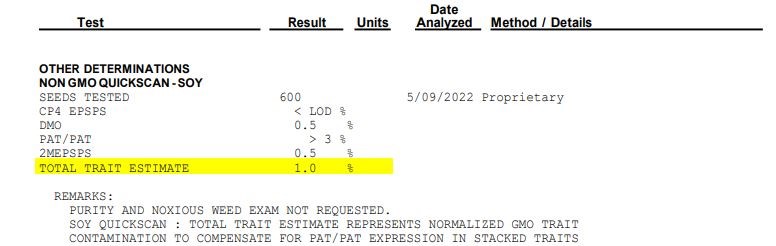
Please note that if PAT/pat from multiple trait stacks is detected, the total trait estimate will not equal the combined individual contamination levels. The normalization algorithm will adjust the total, but still maintain the level of PAT/pat detected in the sample. When making decisions based on results from this test, it is essential consider the estimated total adjusted for trait stacks.
Please contact Field Services Director, Zach Duray at zduray@ilcrop.com or 217-359-4053 with any questions regarding non-GMO testing.
